Research
We are a group of Analytical Chemists who are passionate about application of Data Science and High Resolution Mass Spectrometry (HRMS) for unravelling the human and environmental exposome.
We build digital Machine Learning (ML) based tools to process LC/GC-HRMS data generated from complex samples from environmental to biological. These tools are to automize the data processing from the data import to structural elucidation and the assessment of quality of the identification. Additionally, we build models to maximize the level of information extracted from the spectra of structurally unknown chemicals. Our models take advantage the structural information implicitly present in the fragmentation pattern of an organic chemical. These complex models use a combination of chromatographic and mass spectrometric behavior of organic chemicals to infer about their environmental fate and toxicity.
Open Science
All our tools are built following the FAIR Principals and made publicly available through GitHub and Bitbucket with MIT license.
Relevant Publications
Updates
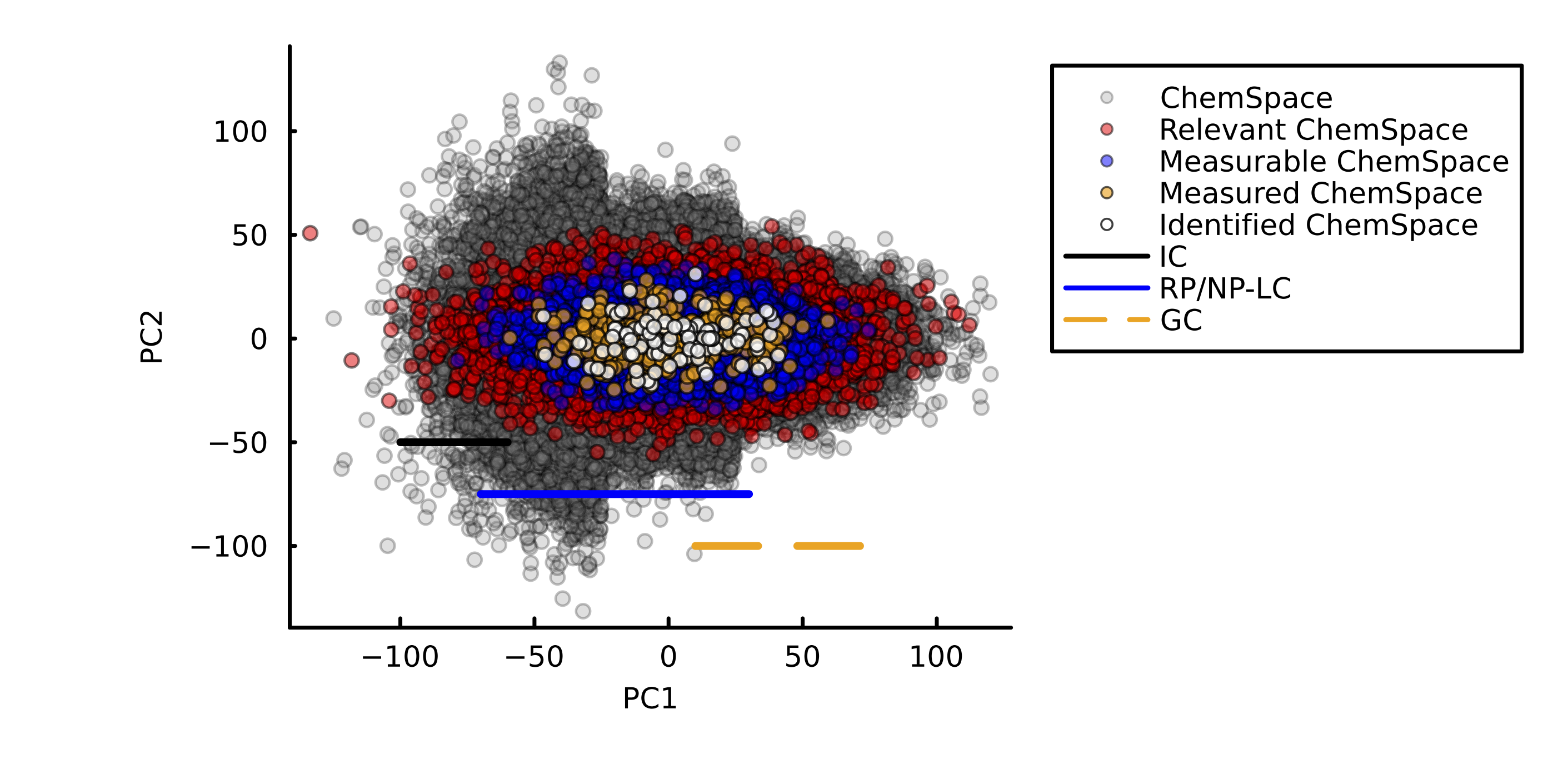
New Paper - Method specific chemical space coverage - March 2, 2026
New article - Tox-based prioritization of unknown features - April 1, 2025
New Publication - Distribution of NPS in wastewater - April 1, 2024
New Preprint - Exploration of Chemical Space - March 11, 2024
New Preprint - Optimization of Molecular Fingerprints - March 1, 2024
